Human mtDNA paper accepted by NAR
Our new paper regarding rNMP incorporation characteristics in human mitochondrial DNA is accepted by Nucleic Acids Research
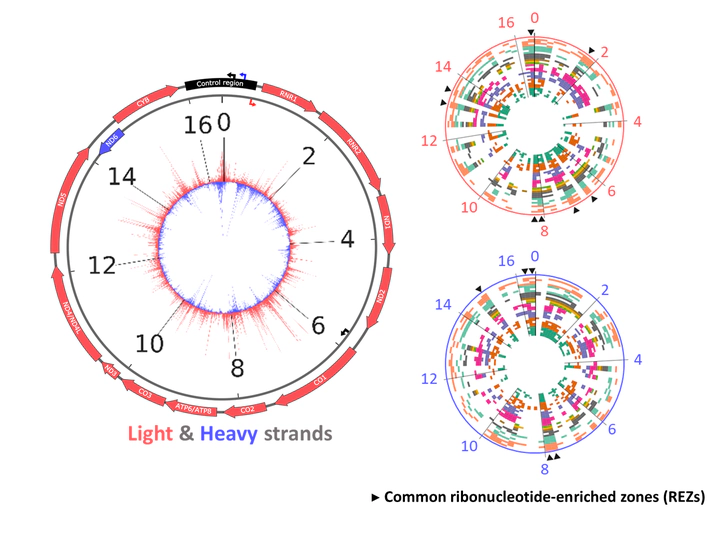
Our new paper, titled “Light-strand bias and enriched zones of embedded ribonucleotides are associated with DNA replication and transcription in the human-mitochondrial genome” is accepted in Nucleic Acids Research. In this paper, we revealed the rNMP incorporation characteristics, and its relation to replication and protein coding genes in human mtDNA. We hope this paper can help us to uncover some functionality of the abundant rNMP incorporation in DNA.
Thanks Dr. Francesca Storici for all the guidance and support in this study. Thanks my co-first author, Taehwan Yang for his tremendous effort and input to this paper. Thanks all the authors for your help and contribution: Kundnani, Deepali; Sun, Mo; Marsili, Stefania; Gombolay, Alli; Jeon, Youngkyu; Newnam, Gary; Balachander, Sathya; Bazzani, Veronica; Baccarani, Umberto; Park, Vivian; Tao, Sijia; Lori, Adriana; Schinazi, Raymond; Kim, Baek; Pursell, Zachary; Tell, Gianluca; Vascotto, Carlo
Here is the full abstract of this paper. I’m always willing to talk if you have any questions or ideas about our study!
Abundant ribonucleoside-triphosphate (rNTP) incorporation into DNA by DNA polymerases in the form of ribonucleoside monophosphates (rNMPs) is a widespread phenomenon in nature, resulting in DNA-structural change and genome instability. The rNMP distribution, characteristics, hotspots, and association with DNA metabolic processes in human mitochondrial DNA (hmtDNA) remain mostly unknown. Here, we utilize the ribose-seq technique to capture embedded rNMPs in hmtDNA of six different cell types. In most cell types, the rNMPs are preferentially embedded on the light strand of hmtDNA with a strong bias towards rCMPs; while in the liver-tissue cells, the rNMPs are predominately found on the heavy strand. We uncover common rNMP hotspots and conserved rNMP-enriched zones across the entire hmtDNA, including in the control region, which links the rNMP presence to the frequent hmtDNA replication-failure events. We show a strong correlation between coding-sequence size and rNMP-embedment frequency per nucleotide on the non-template, light strand in all cell types, supporting the presence of transient RNA-DNA hybrids preceding light-strand replication. Moreover, we detect rNMP-embedment patterns that are only partly conserved across the different cell types and are distinct from those found in yeast mtDNA. The study opens new research directions to understand the biology of hmtDNA and genomic rNMPs.